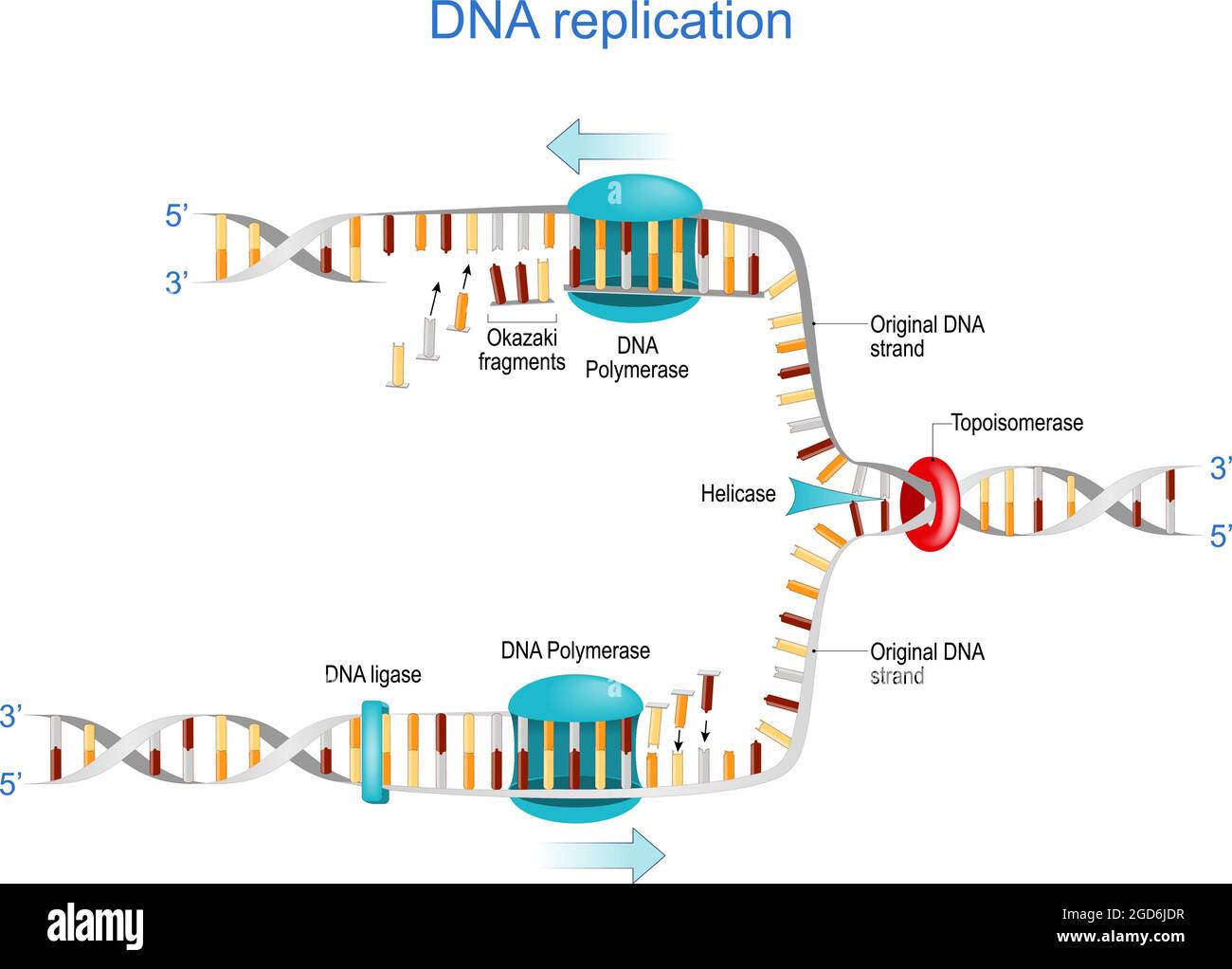
Electron micrograph of Crithidia fasciculata kinetoplast DNA prepared... | Download Scientific Diagram

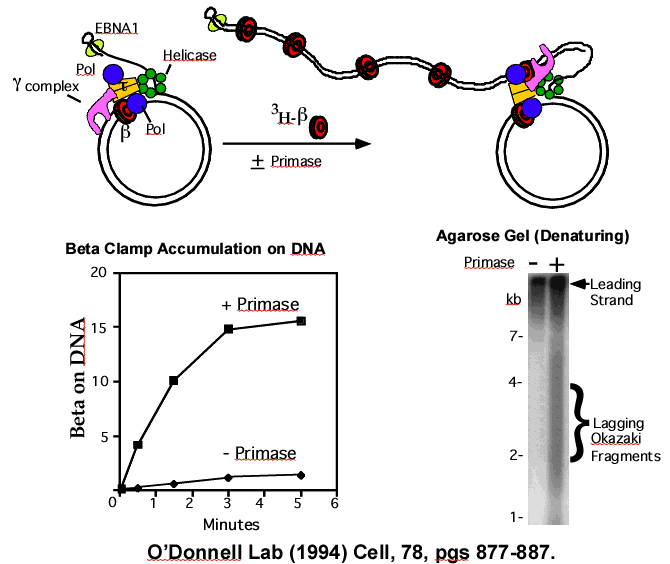
Lecture 1: Fidelity/Specificity: bioregulation through substrate control of molecular choice Use of biochemistry (assays) and genetics (phenotypes) to. - ppt download

Okazaki Fragment Processing-independent Role for Human Dna2 Enzyme during DNA Replication* - Journal of Biological Chemistry

polB completes Okazaki fragment maturation after polD synthesis stops.... | Download Scientific Diagram

Single‐molecule analysis of the Escherichia coli replisome and use of clamps to bypass replication barriers - Georgescu - 2010 - FEBS Letters - Wiley Online Library
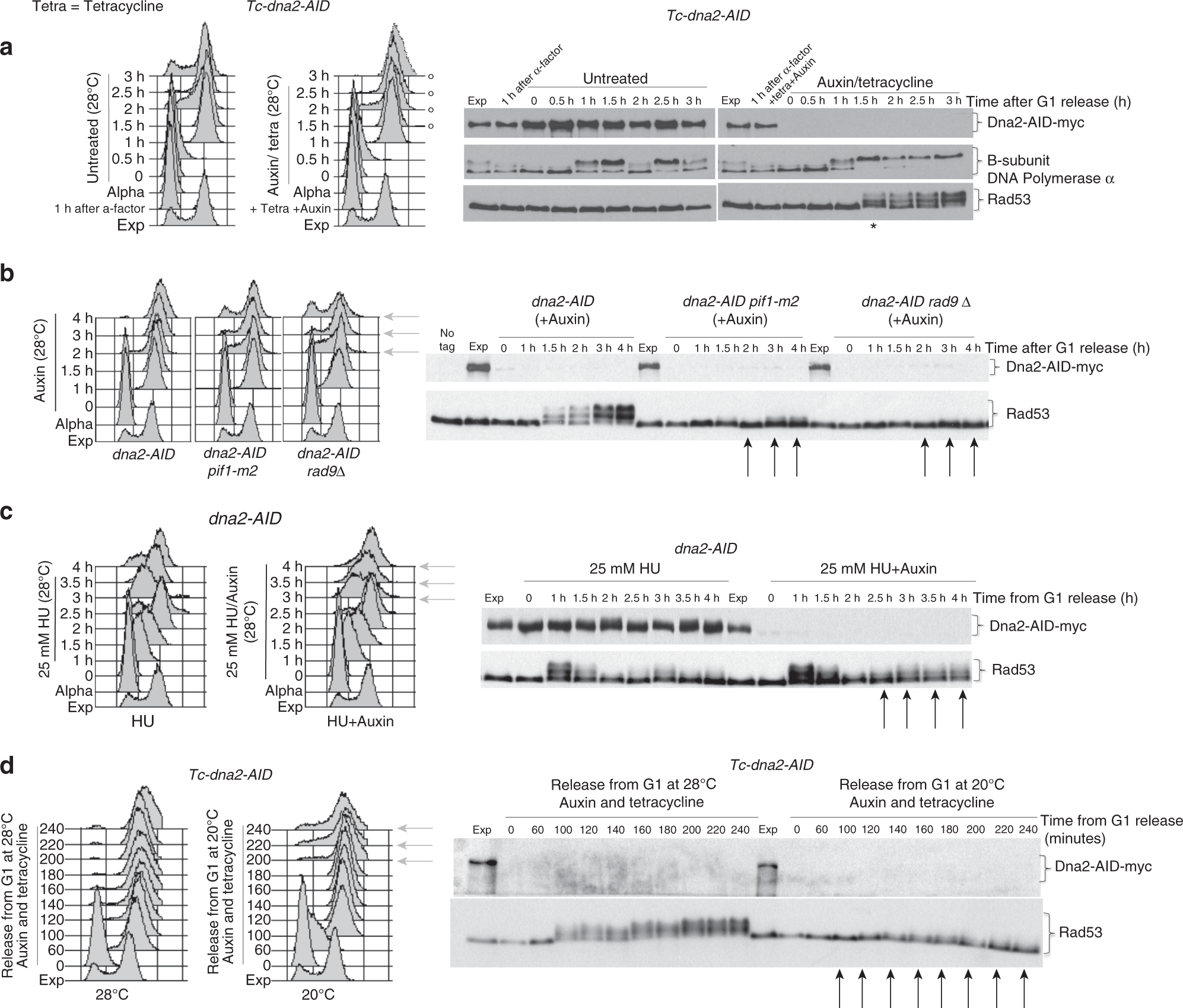
Dna2 processes behind the fork long ssDNA flaps generated by Pif1 and replication-dependent strand displacement | Nature Communications

Analysis of the Okazaki Fragment Distributions along Single Long DNAs Replicated by the Bacteriophage T4 Proteins - ScienceDirect

Analysis of ClaI-cleaved RIs of pSCM201 by electron microscopy. (A and... | Download Scientific Diagram
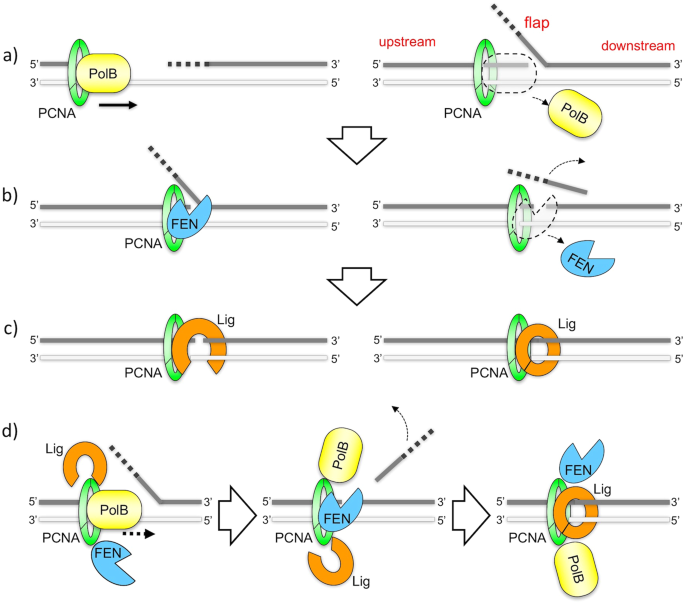
Direct visualization of DNA baton pass between replication factors bound to PCNA | Scientific Reports

The Importance of Poly(ADP-Ribose) Polymerase as a Sensor of Unligated Okazaki Fragments during DNA Replication. - Abstract - Europe PMC

Phosphorylation of PNKP mediated by CDKs promotes end-processing of Okazaki fragments during DNA replication | bioRxiv

Okazaki fragment maturation involves α‐segment error editing by the mammalian FEN1/MutSα functional complex | The EMBO Journal

Okazaki fragment maturation involves α‐segment error editing by the mammalian FEN1/MutSα functional complex | The EMBO Journal